吉林大学学报(工学版) ›› 2025, Vol. 55 ›› Issue (8): 2693-2702.doi: 10.13229/j.cnki.jdxbgxb.20231262
• 计算机科学与技术 • 上一篇
基于多模态数据融合的胃癌患者生存预测模型
刘元宁1,2( ),王星喆1,2,黄子彧3,张家晨1(
),王星喆1,2,黄子彧3,张家晨1( ),刘震1,4
),刘震1,4
- 1.吉林大学 计算机科学与技术学院,长春 130012
2.吉林大学 符号计算与知识工程教育部重点实验室,长春 130012
3.北京林业大学 信息学院,北京 100083
4.长崎综合科学大学 研究生院工学研究科,长崎 851-0193
Stomach cancer survival prediction model based on multimodal data fusion
Yuan-ning LIU1,2( ),Xing-zhe WANG1,2,Zi-yu HUANG3,Jia-chen ZHANG1(
),Xing-zhe WANG1,2,Zi-yu HUANG3,Jia-chen ZHANG1( ),Zhen LIU1,4
),Zhen LIU1,4
- 1.College of Computer Science and Technology,Jilin University,Changchun 130012,China
2.Key Laboratory of Symbolic Computation and Knowledge Engineering of Ministry of Education,Jilin University,Changchun 130012,China
3.School of Information Science and Technology,Beijing Forestry University,Beijing 100083,China
4.Graduate School of Engineering,Nagasaki Institute of Applied Science,Nagasaki 851-0193,Japan
摘要:
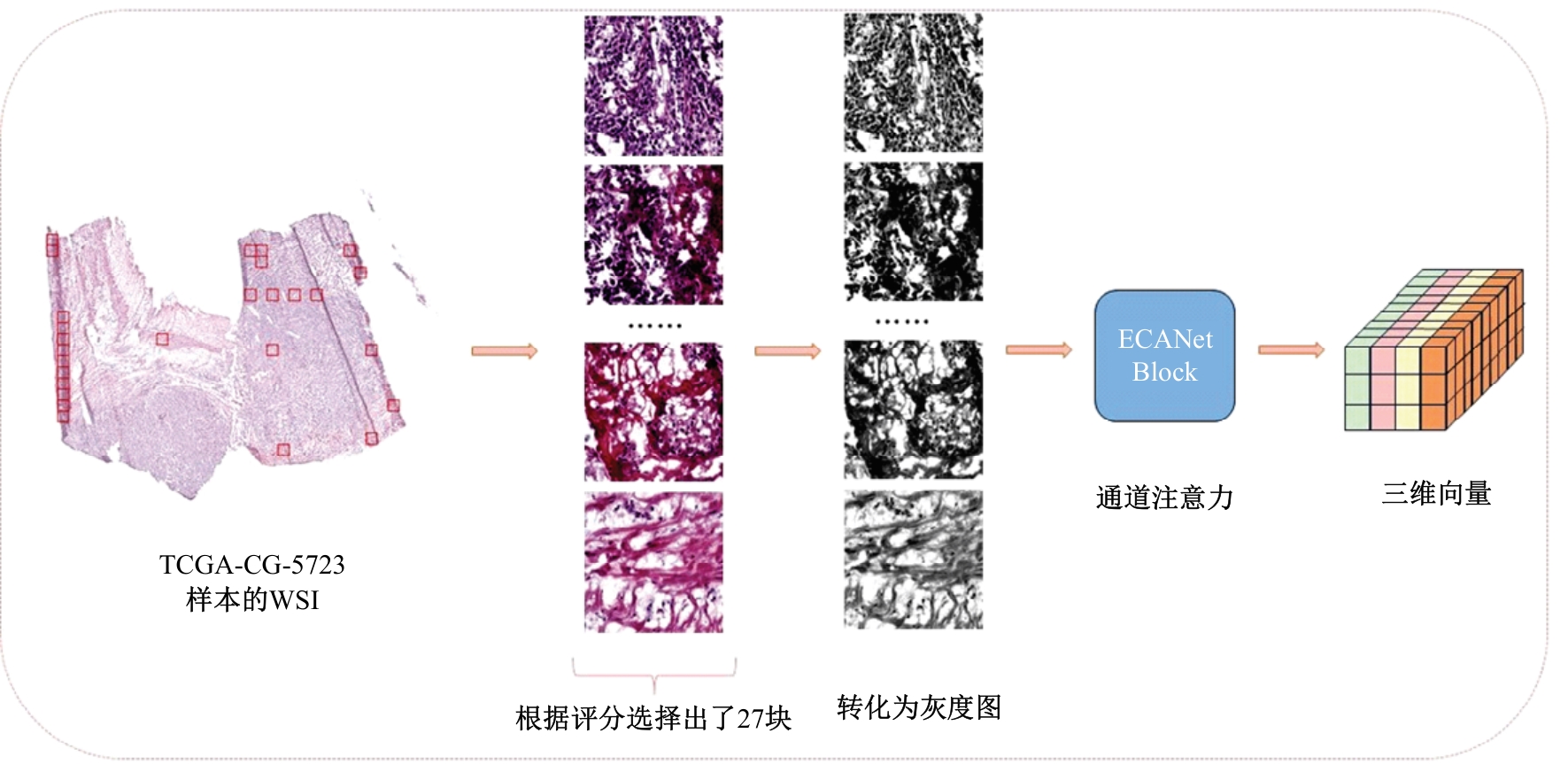
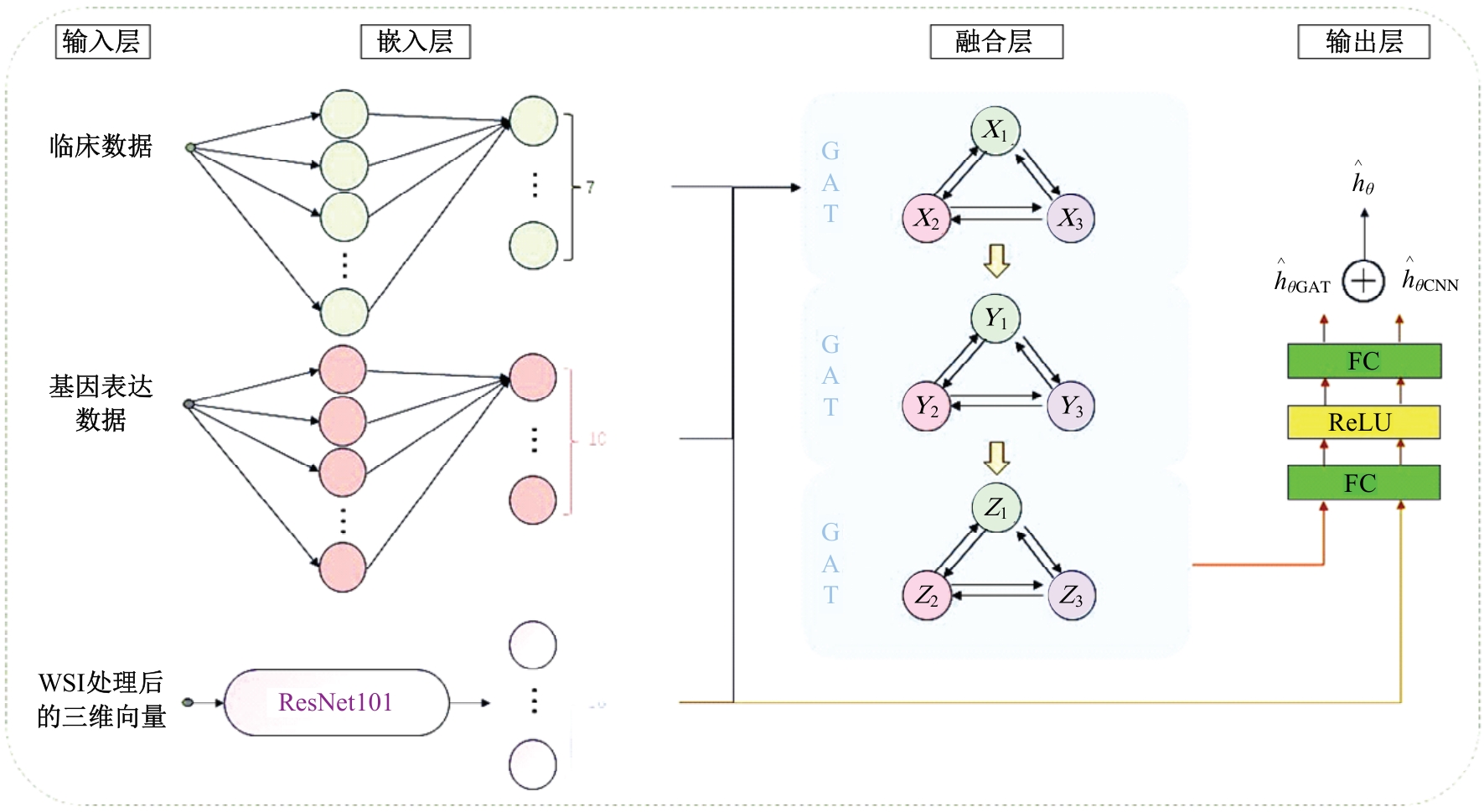
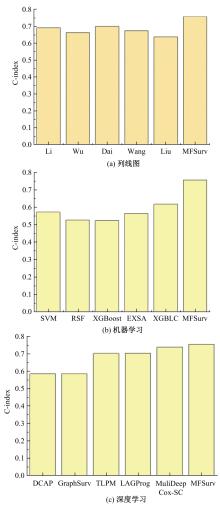
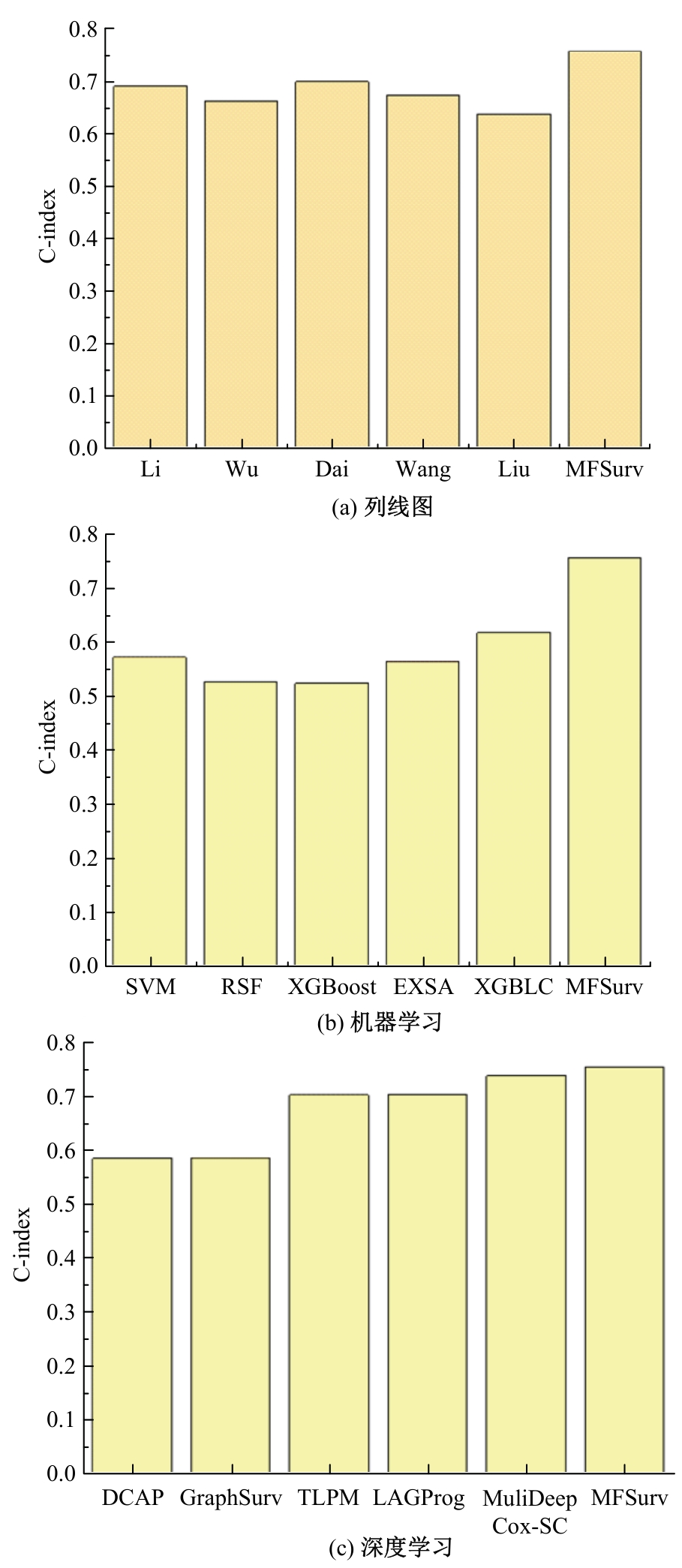
针对胃癌患者生存预测的深度学习方法中存在患者数据使用不全面、数据结合方式粗糙等问题,提出一种多模态数据融合的胃癌患者生存预测模型。首先,将同一患者包括临床数据、基因表达数据和医学图像的多模态数据预处理。其次,将多模态数据输入图注意力网络(GAT)中,使多模态数据在注意力机制下自适应调整权重互相融合。再次,引入卷积神经网络处理的医学图像,与图注意力网络的输出共同作用于预测结果。最后,使用十折交叉验证证明模型性能的稳定性,并将结果与使用相同数据集的其他方法进行比较。实验结果表明,本文提出的模型取得了优秀的准确率。
中图分类号:
- TP391
| [1] | Huang S, Yang J, Fong S, et al. Artificial intelligence in cancer diagnosis and prognosis: Opportunities and challenges[J]. Cancer Letters, 2020, 471: 61-71. |
| [2] | Sung H, Ferlay J, Siegel R L, et al. Global cancer statistics 2020: globocan estimates of incidence and mortality worldwide for 36 cancers in 185 countries[J]. CA: A Cancer Journal for Clinicians, 2021, 71(3): 209-249. |
| [3] | Guan W L, He Y, Xu R H. Gastric cancer treatment: recent progress and future perspectives[J]. Journal of Hematology & Oncology, 2023, 16(1): 57-84. |
| [4] | 覃丽粒, 马小波, 赵天业, 等. MMP-9和TIMP-1表达在胃癌根治术后患者预后评估中的作用[J]. 吉林大学学报: 医学版, 2022, 48(1): 163-171. |
| Qin Li-li, Ma Xiao-bo, Zhao Tian-ye, et al. Effects of MMP-9 and TIMP-1 expressions on prognostic evaluation of gastric cancer patients after radical gastrectomy[J]. Journal of Jilin University (Medicine Edition), 2022, 48(1): 163-171. | |
| [5] | 崔海康, 张旭东, 李晓宁, 等. 细胞焦亡分型和APOD 预测胃癌患者预后作用的生物信息学分析[J]. 吉林大学学报: 医学版, 2023, 49(5): 1268-1279. |
| Cui Hai-kang, Zhang Xu-dong, Li Xiao-ning, et al. Bioinformatics analysis on predition effect of subtypes of cell pyroptosis and APOD on prognosis of gastric cancer patients[J]. Journal of Jilin University (Medicine Edition), 2023, 49(5): 1268-1279. | |
| [6] | Boorn H G, Engelhardt E G, Kleef J, et al. Prediction models for patients with esophageal or gastric cancer: a systematic review and meta-analysis[J]. Plos One, 2018, 13(2): No.e0192310. |
| [7] | Hosny A, Parmar C, Quackenbush J, et al. Artificial intelligence in radiology[J]. Springer Science and Business Media LLC, 2018, 18: 500-510. |
| [8] | Tran K A, Kondrashova O, Bradley A, et al. Deep learning in cancer diagnosis, prognosis and treatment selection[J]. Genome Medicine, 2021, 13(1): 152-168. |
| [9] | Krizhevsky A, Sutskever I, Hinton G E. ImageNet classification with deep convolutional neural networks[J]. Communications of the ACM, 2017, 60(6): 84-90. |
| [10] | Lecun Y, Bengio Y, Hinton G. Deep learning[J]. Nature, 2015, 521: 436-444. |
| [11] | Katzman J L, Shaham U, Cloninger A, et al. Deepsurv: personalized treatment recommender system using a cox proportional hazards deep neural network[J]. BMC Medical Research Methodology, 2018, 18(1): 24-35. |
| [12] | Choi S, Kim S. Artificial intelligence in the pathology of gastric cancer[J]. Journal of Gastric Cancer, 2023, 23(3): 410-427. |
| [13] | Deepa P, Gunavathi C. A systematic review on machine learning and deep learning techniques in cancer survival prediction[J]. Progress in Biophysics and Molecular Biology, 2022, 174: 62-71. |
| [14] | Lipkova J, Chen R J, Chen B, et al. Artificial intelligence for multimodal data integration in oncology[J]. Cancer Cell, 2022, 40(10): 1095-1110. |
| [15] | Boehm K M, Khosravi P, Vanguri R, et al. Harnessing multimodal data integration to advance precision oncology[J]. Nature Reviews Cancer, 2021, 22(2): 114-126. |
| [16] | Wei T, Yuan X, Gao R, et al. Survival prediction of stomach cancer using expression data and deep learning models with histopathological images[J]. Cancer Science, 2022, 114(2): 690-701. |
| [17] | Cox D R. Regression models and life-tables[J]. Journal of the Royal Statistical Society, Series B (Statistical Methodology), 1972, 34(2): 187-202. |
| [18] | Cox D R, Oakes D. Analysis of survival data[J]. Biometrics, 1985, 41(2): 593-594. |
| [19] | Tarkhan A, Simon N, Bengtsson T, et al. Survival prediction using deep learning[C]∥Proceedings of AAAI Spring Symposium on Survival Prediction–Algorithms, Challenges, and Applications, Virtual, USA, 2021: 1-8. |
| [20] | Li H, Lin D, Yu Z, et al. A nomogram model based on the number of examined lymph nodes–related signature to predict prognosis and guide clinical therapy in gastric cancer[J]. Frontiers in Immunology, 2022, 13: No.947802. |
| [21] | Wu J, Wang X, Wang N, et al. Identification of novel antioxidant gene signature to predict the prognosis of patients with gastric cancer[J]. World Journal of Surgical Oncology, 2021, 19(1): No.1901219. |
| [22] | Dai W, Xiao Y, Tang W, et al. Identification of an emt-related gene signature for predicting overall survival in gastric cancer[J]. Frontiers in Genetics, 2021, 12: No. 661306. |
| [23] | Wang M, Jing J, Li H, et al. The expression characteristics and prognostic roles of autophagy-related genes in gastric cancer[J]. PeerJ, 2021, 9: No.e10814. |
| [24] | Liu M, Li J, Huang Z, et al. Gastric cancer risk-scoring system based on analysis of a competing endogenous RNA network[J]. Translational Cancer Research, 2020, 9(6): 3889-3902. |
| [25] | Kourou K, Exarchos T P, Exarchos K P, et al. Machine learning applications in cancer prognosis and prediction[J]. Computational and Structural Biotechnology Journal, 2015, 13: 8-17. |
| [26] | Shivaswamy P K, Chu W, Jansche M. A support vector approach to censored targets[C]∥ Proceedings of Seventh IEEE International Conference on Data Mining, Omaha, USA, 2007: 655-660. |
| [27] | Ishwaran H, Kogalur U B, Blackstone E H, et al. Random survival forests[J]. Annals of Applied Statistics, 2008, 2: No.AOAS169. |
| [28] | Chen T, Guestrin C. XGBoost: a scalable tree boosting system[C]∥Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Francisco, USA, 2016: 785-794. |
| [29] | Liu P, Fu B, Yang S X, et al. Optimizing survival analysis of xgboost for ties to predict disease progression of breast cancer[J]. Institute of Electrical and Electronics Engineers, 2021, 68: 148-160. |
| [30] | Ma B, Yan G, Chai B, et al. XGBLC: an improved survival prediction model based on XGBoost[J]. Bioinformatics, 2021, 38(2): 410-418. |
| [31] | Li G, Huo D, Guo N, et al. Integrating multiple machine learning algorithms for prognostic prediction of gastric cancer based on immune-related lncRNAs[J]. Frontiers in Genetics, 2023, 14: No.1106724. |
| [32] | Chai H, Zhou X, Zhang Z, et al. Integrating multi-omics data through deep learning for accurate cancer prognosis prediction[J]. Computers in Biology and Medicine, 2021, 134: No.104481. |
| [33] | Wang Y, Zhang Z, Chai H, et al. Multi-omics cancer prognosis analysis based on graph convolution network[C]∥Proceedings of IEEE International Conference on Bioinformatics and Biomedicine, Houston, USA, 2021: 1564-1568. |
| [34] | Zhang Y, Xiong S, Wang Z, et al. Local augmented graph neural network for multi-omics cancer prognosis prediction and analysis[J]. Methods, 2023, 213: 1-9. |
| [35] | Avelar P H C, Tavares A R, Silveira T L T, et al. Superpixel image classification with graph attention networks[C]∥Proceedings of 33rd SIBGRAPI Conference on Graphics, Patterns and Images, Porto de Galinhas, Brazil, 2020: 203-209. |
| [36] | Velickovic P, Cucurull G, Casanova A, et al. Graph Attention Networks[EB/OL]. [2023-11-04]. . |
| [37] | Fu X, Patrick E, Yang J Y H, et al. Deep multimodal graph-based network for survival prediction from highly multiplexed images and patient variables[J]. Computers in Biology and Medicine, 2023, 154: No.106576. |
| [38] | Duan M, Wang Y, Zhao D, et al. Orchestrating information across tissues via a novel multitask GAT framework to improve quantitative gene regulation relation modeling for survival analysis[J]. Briefings in Bioinformatics, 2023, 24(4): 1-10. |
| [39] | Ye L, Zhang Y, Yang X, et al. An ovarian cancer susceptible gene prediction method based on deep learning methods[J]. Frontiers in Cell and Developmental Biology, 2021, 9: No.730475. |
| [40] | Adeoye J, Hui L, Koohi M M, et al. Comparison of time-to-event machine learning models in predicting oral cavity cancer prognosis[J]. International Journal of Medical Informatics, 2022, 157: No.104635. |
| [41] | Lerademacher J, Wang X. Time-to-event data: an overview and analysis considerations[J]. Journal of Thoracic Oncology, 2021, 16(7): 1067-1074. |
| [42] | Amin M B, Greene F L, Edge S B, et al. The eighth edition AJCC cancer staging manual: continuing to build a bridge from a population-based to a more "personalized" approach to cancer staging[J]. CA-A Cancer Journal for Clinicians, 2017, 67(2): 93-99. |
| [43] | Marcolini A, Bussola N, Arbitrio E, et al. Histolab: a python library for reproducible digital pathology preprocessing with automated testing[J]. Software X, 2020, 20: No.101237. |
| [44] | He K, Zhang X, Ren S, et al. Deep residual learning for image recognition[C]∥Proceedings of 2016 IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, USA, 2016: 770-778. |
| [45] | Brentnall A R, Cuzick J. Use of the concordance index for predictors of censored survival data[J]. Statistical Methods in Medical Research, 2018, 27(8): 2359-2373. |
| [46] | Kaplan E L, Meier P. Nonparametric estimation from incomplete observations[J]. Journal of the American Statistical Association, 1958, 53(282): 457-481. |
| [47] | Pan Q K, Xu X X, Qi C, et al. Feature fusion: graph attention network and cnn combing for hyperspectral image classification[C]∥Proceedings of the 5th International Conference on Control and Computer Vision, New York, USA, 2022: 171-178. |
| [48] | Xie Y, Niu G, Da Q, et al. Survival prediction for gastric cancer via multimodal learning of whole slide images and gene expression[C]∥Proceedings of 2022 IEEE International Conference on Bioinformatics and Biomedicine, Las Vegas, USA, 2022: 1311-1316. |
| [49] | 张德洪, 郑明珠, 李家秋, 等. 基于MSR1 mRNA 和蛋白在泛癌组织中表达的生物信息学分析及其意义[J]. 吉林大学学报: 医学版, 2023, 49(2): 425-439. |
| Zhang De-hong, Zheng Ming-zhu, Li Jia-qiu, et al. Bioinformatics analysis based on expressions of MSR1 mRNA and protein in pan-cancer tissue and its significance[J]. Journal of Jilin University (Medicine Edition), 2023, 49(2): 425-439. | |
| [50] | Selvaraju R R, Cogswell M, Das A, et al. Grad-CAM:visual explanations from deep networks via gradient-based localization[C]∥2017 IEEE International Conference on Computer Vision, Venice, Italy, 2017: 618-626. |
| [51] | Krzyziński M, Spytek M, Baniecki H, et al. Survshap(t): time-dependent explanations of machine learning survival models[J]. Knowledge-Based Systems, 2023, 262: 110234. |
| [1] | 袁靖舒,李武,赵兴雨,袁满. 基于BERTGAT-Contrastive的语义匹配模型[J]. 吉林大学学报(工学版), 2025, 55(7): 2383-2392. |
| [2] | 徐慧智,郝东升,徐小婷,蒋时森. 基于深度学习的高速公路小目标检测算法[J]. 吉林大学学报(工学版), 2025, 55(6): 2003-2014. |
| [3] | 张汝波,常世淇,张天一. 基于深度学习的图像信息隐藏方法综述[J]. 吉林大学学报(工学版), 2025, 55(5): 1497-1515. |
| [4] | 李健,刘欢,李艳秋,王海瑞,关路,廖昌义. 基于THGS算法优化ResNet-18模型的图像识别[J]. 吉林大学学报(工学版), 2025, 55(5): 1629-1637. |
| [5] | 文斌,丁弈夫,杨超,沈艳军,李辉. 基于自选择架构网络的交通标志分类算法[J]. 吉林大学学报(工学版), 2025, 55(5): 1705-1713. |
| [6] | 李振江,万利,周世睿,陶楚青,魏巍. 基于时空Transformer网络的隧道交通运行风险动态辨识方法[J]. 吉林大学学报(工学版), 2025, 55(4): 1336-1345. |
| [7] | 赵孟雪,车翔玖,徐欢,刘全乐. 基于先验知识优化的医学图像候选区域生成方法[J]. 吉林大学学报(工学版), 2025, 55(2): 722-730. |
| [8] | 徐慧智,蒋时森,王秀青,陈爽. 基于深度学习的车载图像车辆目标检测和测距[J]. 吉林大学学报(工学版), 2025, 55(1): 185-197. |
| [9] | 刘元宁,臧子楠,张浩,刘震. 基于深度学习的核糖核酸二级结构预测方法[J]. 吉林大学学报(工学版), 2025, 55(1): 297-306. |
| [10] | 张磊,焦晶,李勃昕,周延杰. 融合机器学习和深度学习的大容量半结构化数据抽取算法[J]. 吉林大学学报(工学版), 2024, 54(9): 2631-2637. |
| [11] | 李路,宋均琦,朱明,谭鹤群,周玉凡,孙超奇,周铖钰. 基于RGHS图像增强和改进YOLOv5网络的黄颡鱼目标提取[J]. 吉林大学学报(工学版), 2024, 54(9): 2638-2645. |
| [12] | 郭昕刚,何颖晨,程超. 抗噪声的分步式图像超分辨率重构算法[J]. 吉林大学学报(工学版), 2024, 54(7): 2063-2071. |
| [13] | 乔百友,武彤,杨璐,蒋有文. 一种基于BiGRU和胶囊网络的文本情感分析方法[J]. 吉林大学学报(工学版), 2024, 54(7): 2026-2037. |
| [14] | 张丽平,刘斌毓,李松,郝忠孝. 基于稀疏多头自注意力的轨迹kNN查询方法[J]. 吉林大学学报(工学版), 2024, 54(6): 1756-1766. |
| [15] | 孙铭会,薛浩,金玉波,曲卫东,秦贵和. 联合时空注意力的视频显著性预测[J]. 吉林大学学报(工学版), 2024, 54(6): 1767-1776. |
|
||